SNPrank[1] is an eigenvector centrality algorithm that ranks the importance of single nucleotide polymorphisms (SNPs) in a genetic association interaction network (GAIN) [2]. Each SNP is ranked according to its overall contribution to the phenotype, including its main effect and second- and higher-order gene-gene interactions. SNPrank is open source software and available as a command-line tool in several language implementations.
For access to command-line implementations of SNPrank (Matlab, Python, Java, as well as CPU and GPU versions), see our organization page at github.
GAIN can be combined with SNPrank for a powerful analysis engine. We have a tutorial describing the steps of the analysis as well as the dependencies required.
References
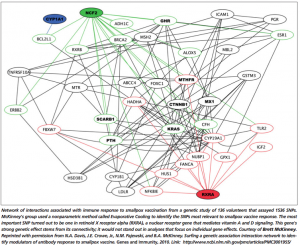
[1]N.A. Davis, J.E. Crowe, Jr., N.M. Pajewski, and B.A. McKinney.
Surfing a genetic association interaction network to identify
modulators of antibody response to smallpox vaccine. Genes and
Immunity, 2010, doi: 10.1038/gene.2010.3. (open
access)
[2]B.A. McKinney, J.Guo, J.E. Crowe, Jr., and D. Tian. Capturing the
spectrum of interaction effects in genetic association studies by
simulated evaporative cooling network analysis. PLoS Genetics 2009,
5(3): e1000432. doi:10.1371/journal.pgen.1000432. (open
access)